In Homework#7 of the 2014 fall AML610 course, we discussed an example where we had a time series of data for a disease vector, V, that can spread disease to a human population. Once the humans catch the disease, they recover after 1/gamma days and moved to the recovered compartment.
Our model looks like this
dS/dt = - beta*V*S/N dI/dt = + beta*V*S/N - gamma*I dR/dt = gamma*I
Somewhat uniquely, in this analysis what we observe in the data is people recovering (not people being infected), so the number of new recoveries in each unit time is X=gamma*I.
I’ve decided it might be more realistic, for the disease we examine in that homework, to add an additional human-to-human infection component, because otherwise we assume that all the transmission of the disease comes only from the vector. So here is the new model:
dS/dt = - beta*V*S/N - mu*I*S/N dI/dt = + beta*V*S/N + mu*I*S/N - gamma*I dR/dt = gamma*I
Recall that in Homework #7 I pointed out that we just have a data time series for V, we don’t have an analytic expression for it. This makes it hard, if not impossible, to use the R deSolve libraries to numerically solve our system of equations for S, I, and R.
A solution is to use a simple time differencing method (Euler’s method) to solve the system of equations, first by re-casting them as time-difference equations:
S_i+1 = S_i - delt*beta*V_i*S_i/N - delt*mu*I_i*S_i/N I_i+1 = I_i + delt*beta*V_i*S_i/N + delt*mu*I_i*S_i/N - delt*gamma*I_i R_i+1 = R_i + delt*gamma*I_i
where V_i is the value of V at the i^th time step, and delt is the time step. We start off the simulations with S_1=N, I_1=0, and R_1=0. In the R script model_fit_example_utils.R, I provide a method called solve_model that takes as its arguments V, beta, mu, gamma, delt, and N, and returns the estimated number of newly observed recovered people per unit time, Xpred.
In the file model_fit_example_data.txt I provide the time series of data for V, and X, which is the observed number of people “recovering” each day.
In the R script model_fit_example.R, I read in this data, and then try to fit our SIRV model above to it. I give options to choose the time step, and then I fit a spline to the available V data to interpolate the data within each day, in steps of delt. The script also randomly sweeps the parameters, and calculates the Pearson chi-squared statistic comparing the model to the data.
In the fit, we use the Pearson chi-squared as our goodness-of-fit statistic (which is appropriate because the X are count data). However, the data are probably over-dispersed, and we will need to correct for that to get reliable confidence intervals on our parameters.
If data are appropriate for the application of a Pearson chi-squared fit, a standard method for correction of over- or under-dispersion developed by McCullagh and Nelder is to multiply the Pearson chi-squared statistic, T, by (N-p)/T_best, where T_best is the value of the Pearson chi-squared statistic at the best-fit values of the model parameters, and p is the number of parameters being fit for. We would then determine the uncertainties on the parameters through examination of T’=(N-p)/T_best, rather than T. In this case, we are fitting for three parameters; beta, mu, and gamma.
Doing the same thing in C++
You’ll probably notice that the R script runs slowly if you choose a small time step like delt=0.01. It would thus be much faster to do these sweeps with a program in C++.
A problem that must be overcome is that if you are running programs on a super-computing system, it is not ideal to have those programs reading files to obtain the data. Much simpler is to hard-code your data within your C++ code.
What I always do it write an R script that prints out the data, in a format where it is already in C++ code form (because I have better things to do with my time than spend hours entering data into a C++ program by hand). Then I can just insert those lines of code into my C++ program. In the R script model_fit_example.R, you’ll see there is a part of the script where I use a lot of cat statements to print lines like
_Vinter.push_back(...);
where the lines fill the brackets with my data. Using the sink() command, I print these lines to a file called code.txt
I’ve created a class called DataClass.h and DataClass.cpp, where I insert the lines of code in code.txt into the constructor of the class. The class also has a method SolveModel, that given hypotheses for beta, mu and gamma, it will use Euler’s method to get the predicted values for X.
In the program Modelling_example.cpp, I show how this class is in a random parameter sweep procedure, printing out to the screen the values of the parameters for each sweep, and also the value of the corresponding goodness-of-fit. Notice that I use getpid() in the program to get the process ID of the program to set my random seed. This is because it turns out that it is difficult to pass arguments at run time to programs running on the NSF XSEDE batch system, so I had to find a work-around for my usual method.
The makefile makefile_data compiles the program, linking it to DataClass and the UsefulUtils.h and UsefulUtils.cpp files.
Try running the file on your laptop
We are now ready to submit our first batch job to XSEDE
Login to the Stampede super-computing system
ssh <your username>@stampede.tacc.xsede.org
Create a new directory called model, that we will be using to put our program code:
mkdir model
stay logged in to Stamped for now.
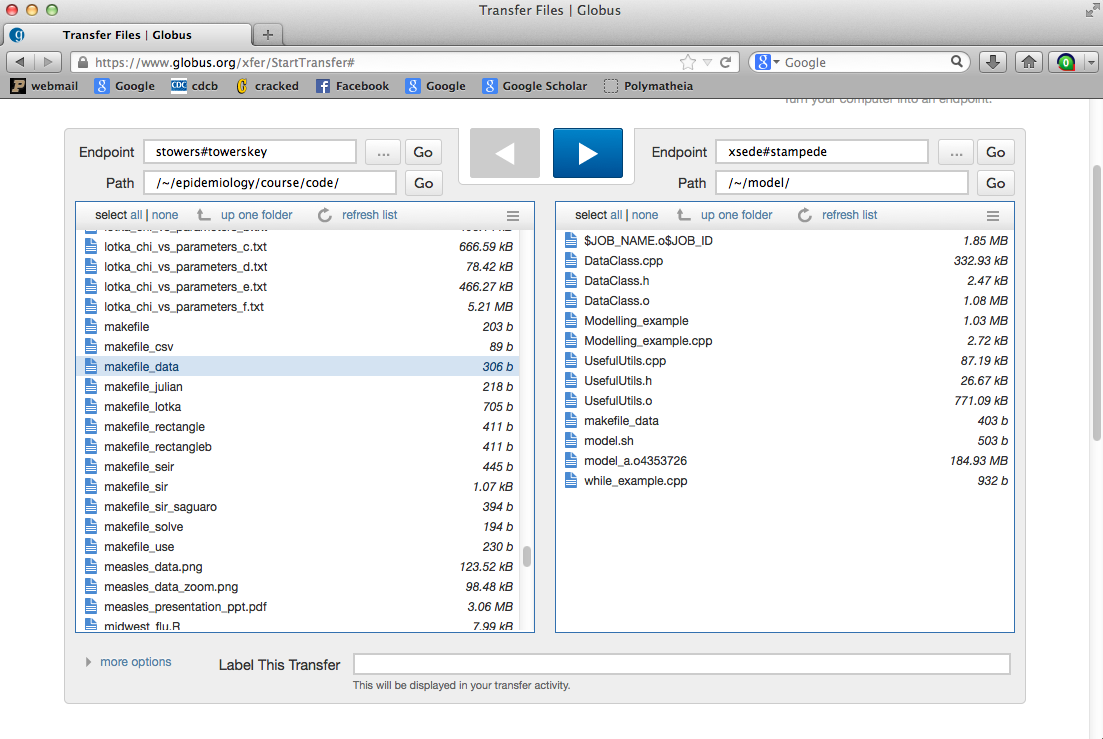
Now start up to Globus file transfer program that you downloaded to your laptop earlier. Copy all the .cpp and .h files associated with our Modelling_example.cpp program.
Here’s what it looked like when I did it:
Now, on Stampede, change to your model directory, and list it:
cd model ls -alrt model
The files should all be there if everything worked correctly.
It turns out that on Stampede they prefer you to use a special C++ compiler called icpc. In the makefile makefile_stampede, I’ve made the modifications such that you can compile the Modelling_example.cpp program on Stampede. Download it to your laptop, then using Globus, copy it over to Stampede too.
To compile your program on Stampede, type
make -f makefile_stampede Modelling_example
It should compile successfully.
Now, it is forbidden to run programs interactively on batch systems (ie; you can lose your account if they catch you doing it). Instead, we have to write what is known as a batch submission script to tell the batch system how many programs we want to run in parallel, how long, maximum, we would like each job to run, where we would like the output directed, etc. In the file model.sh, I’ve put a batch submission script that will run our program on 32 processors on Stampede, for a maximum run time of 10 minutes per job. Download this to your laptop, and then upload it to Stampede using Globus.
To submit this job, type on Stampede
sbatch model.sh
You’ll see a bunch of text like this appear:
----------------------------------------------------------------- Welcome to the Stampede Supercomputer ----------------------------------------------------------------- --> Verifying valid submit host (login2)...OK --> Verifying valid jobname...OK --> Enforcing max jobs per user...OK --> Verifying availability of your home dir (/home1/03096/stowers)...OK --> Verifying availability of your work dir (/work/03096/stowers)...OK --> Verifying availability of your scratch dir (/scratch/03096/stowers)...OK --> Verifying valid ssh keys...OK --> Verifying access to desired queue (development)...OK --> Verifying job request is within current queue limits...OK --> Checking available allocation (TG-DMS140043)...OK Submitted batch job 4356817
The job submitted successfully!
You can check the status of your job by typing
squeue -u <your username>
When I did this, I got
login2.stampede(12)$ squeue -u stowers JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON) 4356817 development model stowers R 0:52 2 c557-[802,901]
If you wait too long to check the queue, your job will have ended, and no output will appear when you use the squeue command.
After my job ended, I listed my directory and found the file model.o4356817 in my directory (yours will be named differently). It contains the output of all the programs that ran in parallel, plus some extra lines at the top that you’ll have to delete. Because the actual file I got was huge, I’ve only included the first several lines for you to look at. You can transfer this file back to your laptop using Globus (but I’d suggest gzipping it first by typing gzip <filename>).
Note that sometimes if Globus has been sitting idle for a while, in order to get the transfer to work you need to “refresh list” on both the local and remote machines.
You have now successfully run your first batch job on a super-computing system!