In this past module, we discussed using the Pearson chi-squared statistic to determine the best-fit parameters of an SIR model to influenza B data from the 2007-08 Midwest flu season. In this module, we will discuss how to find the best-fit parameters using the Negative Binomial likelihood instead.
Category Archives: C++
Submitting jobs to the ASU A2C2 ASURE batch computing system
The AML610 Fall 2014 course has received an allocation of 10,000 CPU hours on the A2C2 Asure batch computing system. Students in the course have received an email from me describing how to sign up for an Asure account under this allocation.
Another example C++ program to fit model parameters to data
In a past module, we examined how we could use methods in the R deSolve to fit the parameters of an SIR model to confirmed cases of influenza B in the Midwest region during the 2007-2008 flu season (the data were obtained from the CDC). In that module, we used a Least Squares goodness-of-fit estimator. Continue reading
A C++ class for numerically solving ODE’s
In previous modules, we have described how to use methods in the R deSolve library to numerically solve systems of ordinary differential equations, like the SIR model. The default algorithm underlying the functions in the deSolve library is 4th order Runge-Kutta method, which involves an iterative process to obtain approximate numerical solutions to the differential equations. Euler’s method is an even simpler method that can be used to estimate solutions to ODE’s, but 4th order Runge-Kutta is a higher order method that is more precise. Continue reading
AML 610 Module XIII: Canadian hare lynx data
Canadian Hare Lynx Data
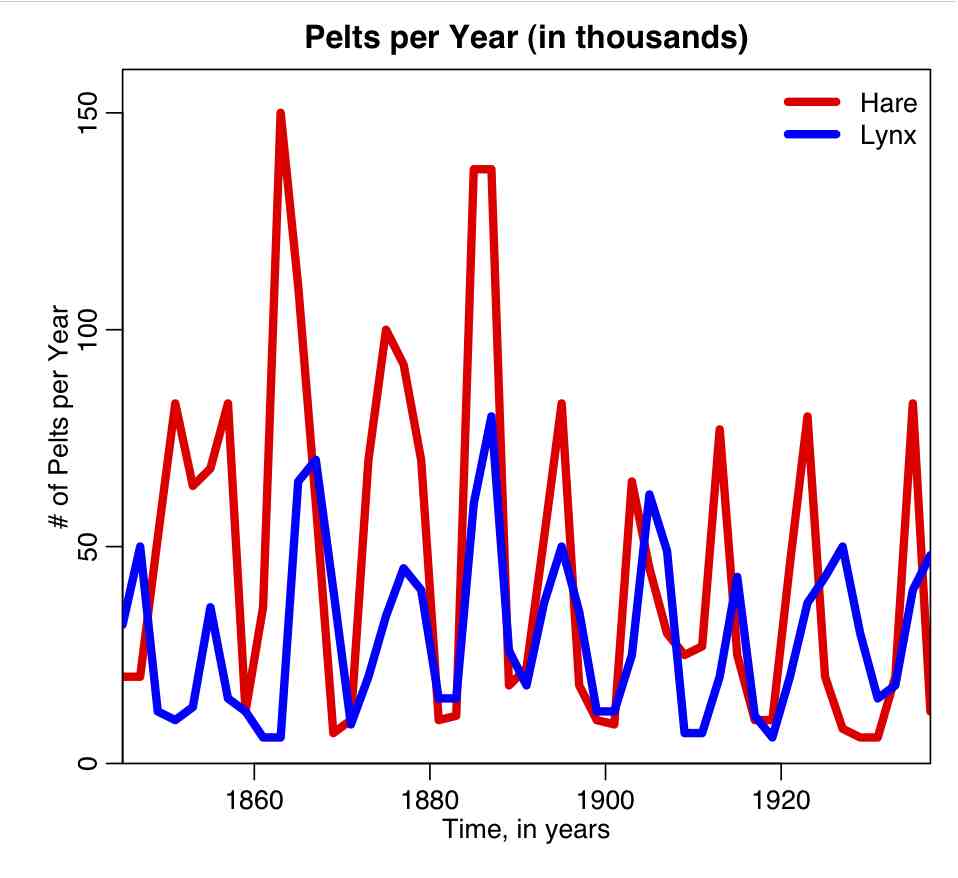
The file hare_lynx.txt contains data on the number of arctic hare and lynx pelts collected by the Hudson’s Bay company in Canada over the course of many years (data obtained from this website). Do you think the Lotka-Volterra model is an appropriate model to fit to these data?
The R script hare_lynx_plot.R plots the Hare Lynx data:
AML 610 Module XII: submitting jobs in batch to the ASU Saguaro distributed-computing system
The ASU Advanced Computing Center (A2C2) maintains the Saguaro distributed computing system, that currently has over 5,000 processor cores.
ASU students in the spring semester of AML610 should have already applied for and received an account on the Saguaro system (per the instructions of last month’s email describing how to apply for an account).
Saguaro allows you to simultaneously run multiple jobs in batch, directing standard output to a log file. For this course, we will be using Saguaro to solve a system of ODE’s under a hypothesis for the parameters and initial conditions values (either chosen in a parameter sweep, or randomly chosen within some range); the output of the ODE’s will then be compared to a data set, and a best-fit statistic (like Least Squares, Pearson chi-squared, or Maximum likelihood) computed. The parameter values and best-fit statistics are then printed to standard output.
Access to cloud computing resources, and knowledge of how to utilize those resources, has many different potential applications in modelling. Learning how to use Saguaro as a tool in solving problems related to this course can thus potentially open up many further avenues of future research to you.
Homework #5, due Thus April 18th, 2013 at 6pm. Data for the homework can be found here.
AML610 module XI: practical problems when connecting deterministic models to data
Some (potentially) useful utilities for random number generation and manipulating vectors in C++
I’ve written some C++ code mainly related to vectors; calculating the weighted mean, running sum, extracting every nth element, etc). There are also utilities related to random number generation from various probability distributions, and methods to calculate the CDF of various probability distributions.
The file UsefulUtils.h and UsefulUtils.cpp contain source code of a class that contains these utilities that can be useful when performing compartmental modelling in C++. These utilities will be used extensively in the examples that will be presented in this, and later, modules. The file example_useful_utils.cpp gives examples of the use of the class. It can be compiled with the makefile makefile_use with the command
make -f makefile_use example_useful_utils
Homework #4, due April 3rd, 2013 at 6pm. The data for the homework can be found here.
Numerical methods to solve ordinary differential equations
After going through this module, students will be familiar with the Euler and Runge-Kutta methods for numerical solution of systems of ordinary differential equations. Examples are provided to show students how complementary R scripts can be written to help debug Runge-Kutta methods implemented in C++.
Contents
- Euler’s method (finite difference method)
- Using the Euler method to solve dN/dt = rho*N
- Implementing the Euler method in C++
- Comparison of the output of the C++ and R programs implementing the Euler method to solve dN/dt=rho*N
- Dynamic time step calculation in the Euler method
- Runge-Kutta method
- Example of implementation of Runge Kutta in C++: Lotka-Volterra predator prey model
ASU AML 610 Module IX: Introduction to C++ for computational epidemiologists
After going through this module, students should be familiar with basic skills in C++ programming, including the structure of a basic program, variable types, scope, functions (and function overloading), control structures, and the standard template library.
- Hello world
- Variable types
- Strings
- Boolean logic
- Scope of variables
- Introduction to functions
- Arguments to main
- Control structures (if/then/else, while, for loops)
- Functions, pass by value
- Functions, pass by reference
- Overloading functions
- The standard template library: vectors
- Vectors in multi-dimensions
- Parsing data from comma delimited files
- Makefiles
- Introduction to classes and objects
So far in this course we have used R to explore methods related to fitting model parameters to data (in particular, we explored the Simplex method for parameter estimation). As we’ve shown, parameter estimation can be a very computationally intensive process.
When you use R, it gives you a prompt, and waits for you to input commands, either directly through the command line, or through an R script that you source. Because R is a non-compiled language, and instead interprets code step-by-step, it does not have the ability to optimize calculations by pre-processing the code.
In contrast, compiled programming languages like C, java, or C++ (to name just a few) use a compiler to process the code, and optimize the computational algorithms. In fact, most compilers have optional arguments related to the level of optimization you desire (with the downside that the optimization process can be computationally intensive). Optimized code runs faster than non-optimized code.
Good programming practices (in any language)
Easy readability, ease of editing, and ease of re-usability are things to strive for in code you write in any language. Achieving that comes with practice, and taking careful note of the hallmarks of tidy, readable, easily editable, and easily re-usable code written by other people.
While I’m certainly not perfect when it comes to utmost diligence in applying good programming practices, I do strive to make readable and re-useable code (if only because it makes my own life a lot easier when I return to code I wrote a year earlier, and I try to figure out what it was doing).
In the basics_programming.R script that I make use of some good programming practices that ensure easy readability. For instance, code blocks that get executed within an if/then statement, for loops, or while loops are indented by a few spaces (usually two or three… be consistent in the number of indent spaces you use). This makes it clear when reading the code which nested block of code you are looking at. I strongly advise you to not use tabs to indent code. To begin with, every piece of code I’ve ever had to modify that had tabs for indent also used spaces here and there for indentation, and it makes it a nightmare to edit and have the code remain easily readable. Also, if you have more than one or two nested blocks of code, using tabs moves the inner blocks too far over to make the code easily readable.
In the R script sir_agent_func.R I define a function. Notice in the script that instead of putting all the function arguments on one long line, I do it like this:
SIR_agent = function(N # population size
,I_0 # initial number infected
,S_0 # initial number susceptible
,gamma # recovery rate in days^{-1}
,R0 # reproduction number
,tbeg # begin time of simulation
,tend # end time of simulation
,delta_t=0 # time step (if 0, then dynamic time step is implemented)
){
This line-by-line argument list makes it really easy to read the arguments (and allows inline descriptive comments). It also makes it really easy to edit, because if you want to delete an argument, it is simple as deleting that line. If you want to add an argument, add a line to the list.
Descriptive variable names are a good idea because they make it easier for someone else to follow your code, and/or make it easier to figure out what your own code did when you look at it 6 months after you wrote it.
Other good programming practices are to heavily comment code, with comment breaks that clearly delineate sections of code that do specific tasks (this makes code easy to read and follow). I like to put such comments “in lights” (ie; I put a long line of comment characters both above and below the comment block, to make it stand out in the code). If the comments are introducing a function, I will usually put two or three long lines of comment characters at the beginning of the comment block; it makes it clear and easy to see when paging through code where the functions are defined.
In-line comments are also very helpful to make it clear what particular lines of code are doing.
A good programming practice that makes code much easier to re-use is to never hard-code numbers in the program. For instance, at the top of the basics_programming.R script I create a vector that is n=1000 elements long. In the subsequent for loop, I don’t have
for (i in 1:1000){}
Instead I have
for (i in 1:n){}
This makes that code reusable as-is if I decide to use it to loop over the elements of a vector with a different length. All I have to do is change n to the length of the new vector. Another example of not hard-coding numbers is found in the code associated with the while loop example.
As an aside here, I should mention that in any programming language you should never hard-code the value of a constant like π (as was pointed out in basics.R, it is a built-in constant in R, so you don’t need to worry about this for R code). In other languages, you should encode pi as pi=acos(-1.0), rather than something like pi=3.14159. I once knew a physics research group that made the disastrous discovery that they had a typo in their hard-coded value of pi… they had to redo a whole bunch of work once the typo was discovered.
Notice in the script that I have a comment block at the top of basics_programming.R that explains what the script does, gives the date of creation and the name of the person who wrote the script (ie; me). It also gives the contact info for the author of the script, and copyrights the script. Every script or program you write should have that kind of boilerplate at the top (even if you think the program you are writing will only be used by you alone… you might unexpectedly end up sharing it with someone, and/or the boilerplate makes it clear that the program is *your* code, and that people just can’t pass it off as their own it if they come across it). It also helps you keep track of when you wrote the program.